Breast Cancer
Biomarker Identified for Likely Aggressive, Early Stage Breast Cancer
The one-size-fits-all approach to early stage breast cancer creates a paradox: Millions of dollars are spent on unnecessary surgeries and radiation to treat women with low-risk ‘in situ’ lesions, an estimated 85% of which would never progress to invasive cancers. Meanwhile, the standard conservative treatment is insufficient for many early-stage tumors that have progressed past the in situ stage and fails to prevent their spread to distant sites in the body.
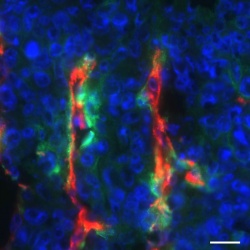
Now Whitehead Institute researchers have identified SMARCE1, a gene overexpressed in the subset of early-stage cancers that are likely to become aggressively invasive – making it possible for the first time to distinguish poorly invasive tumors from those that will likely spread and metastasize. With such a biomarker, doctors could better tailor therapies designed to match the behavior of each patient’s cancer.
The researchers found that 50 percent of the early-stage cancers with high SMARCE1 expression will metastasize at some point in the 10 to 15 years after their initial diagnosis. “Early-stage cancers are not all the same. Some are destined to go rogue and should be treated from the outset with this understanding in mind,” says Whitehead Member Piyush Gupta, who is also an assistant professor of biology at MIT.
Breast cancer begins as anomalous cells that divide out of control, usually in the milk ducts. In almost all cases, a patient does not succumb to the initial cancer but to the secondary tumors after the cancer has spread.
Over the past two decades, mammography’s ability to detect ever more miniscule lesions has significantly increased. As a result, many patients with harmless lesions undergo surgery and radiation therapy. Conversely, current therapies fall short for a quarter of patients with early-stage tumors – which ultimately spread to distant sites in the body. By sorting the aggressive from the benign, doctors could tailor therapies more accurately to each patient and avoid the dual pitfalls of costly overtreatment and potentially fatal undertreatment.
To determine why some lesions are more aggressive than others, scientists in the Gupta lab analyzed the regulators of about 350 genes with increased expression in the invasive regions of cancers. The team, which included then-graduate student Ethan Sokol, postdoctoral researcher Yuxiong Feng, and graduate student Dexter Jin, identified a large group of these genes that allows cancer cells to invade the structural support surrounding the cells, called the extracellular matrix. One gene regulates the group: SMARCE1.
“It’s clear that SMARCE1 is affecting all of the key players in invasion and metastasis,” says Sokol, who along with Feng and Jin authored a paper in PNAS that describes their work. “It’s amazing when you look at the list of the things it’s regulating.”
Interestingly, SMARCE1 only seems to be important during the early stages of metastasis, making it a suitable biomarker for this critical step.
“We looked at every step of the metastatic cascade, and the tumor growth at the primary site, as well as the growth at the distant metastases, are not affected by this gene,” says Feng. “Only the invasion is affected by SMARCE1.”
In fact, when the team analyzed SMARCE1’s activity in a model of human breast tissues created by the Gupta lab, they determined that SMARCE1 is required for localized breast cancer cells to escape into the surrounding tissues. Without it, the cells stay confined and relatively harmless.
The team also analyzed SMARCE1 levels in tissue samples from about 200 early stage breast cancer patients in a retrospective study. Those with the highest SMARCE1 levels were the most likely to suffer from metastasis and have the worst outcomes. The relationship between SMARCE1 levels and prognosis held true for lung and ovarian cancer patients as well, suggesting that the gene’s importance is not limited to one kind of cancer.
In addition to assessing SMARCE1’s role in other forms of cancer, the team is working with oncologists to take the next steps needed to translate their findings to the clinic.
Source: Newswise, written by Nicole Giese Rura
18.04.2017